Just today, the entire Internet was flooded with the story of this post-00s guy! This netizen named Seth Howes used a portable sequencer and Claude to independently complete the complete genome sequencing in his living room, and successfully traced the pathogenic mechanism of autoimmune diseases in his family for multiple generations.
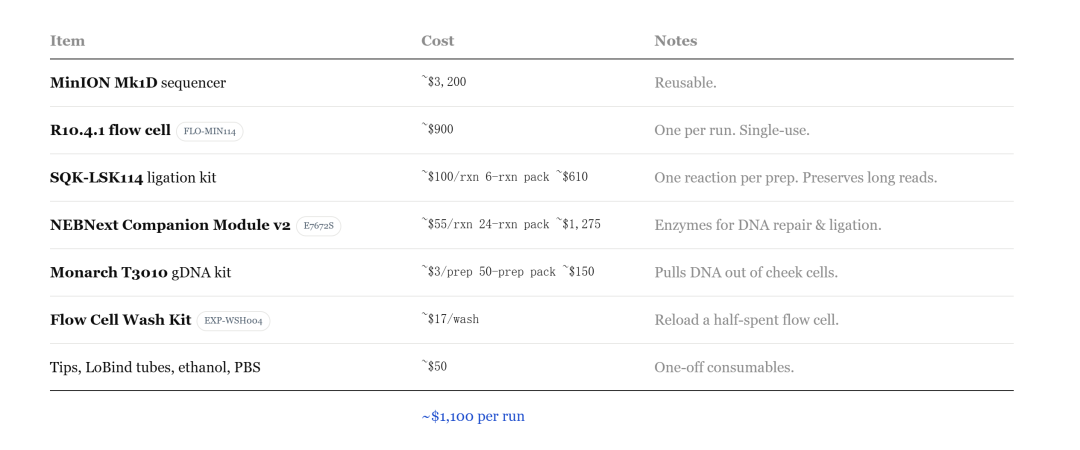
With a sequencer the size of a USB flash drive and several AI models, Seth Howes, a young man born in the 2000s, completed genome sequencing in his living room and single-handedly solved the mystery of his family's autoimmune disease that had remained unsolved for decades. In 2023, the cost of completing a whole human genome sequence will be US$2.7 billion, but he only spent US$1,100!
These problems have left countless clinicians at a loss.
No clinician has been able to explain these mechanisms before. After more than ten years of treatment and countless hospital visits, the answer was finally found in my own living room.
This experiment is so shocking that the biotechnology circle is now in a state of shock!
Claude's large model actually turned gene sequencing from a capital-intensive activity into a personal tool.
The location of biological research is no longer an expensive machine in a top laboratory, nor is it a public platform in a medical school, but in your own living room!
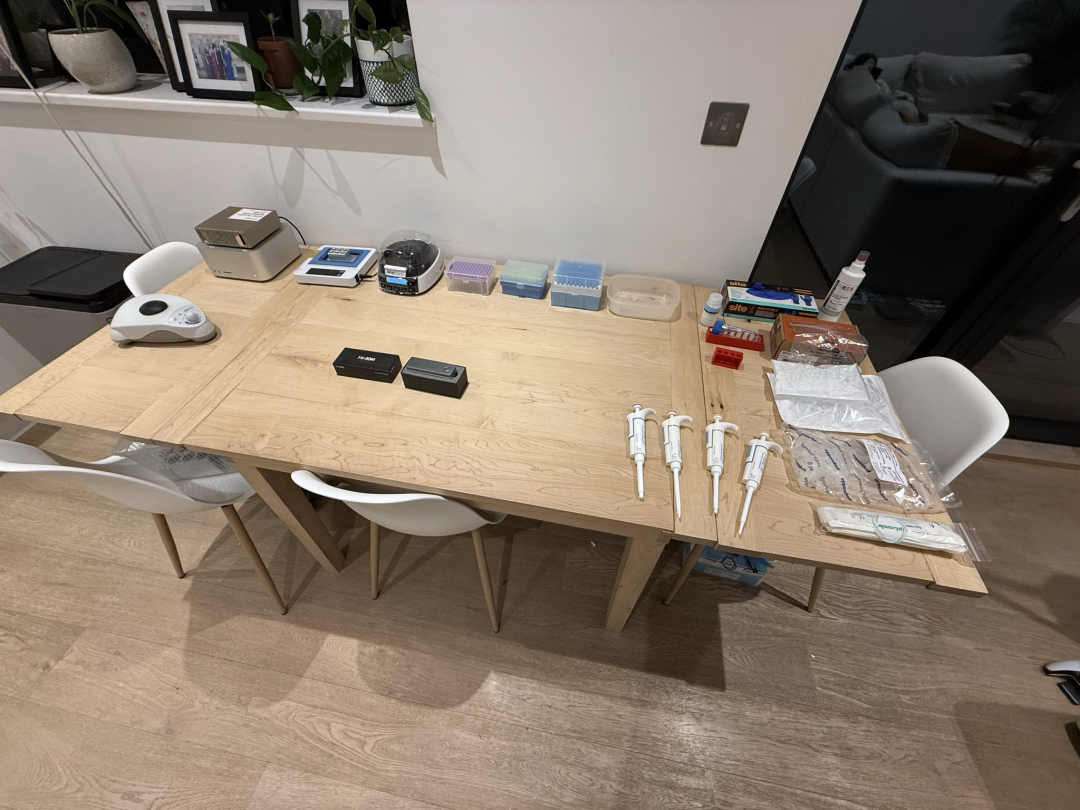

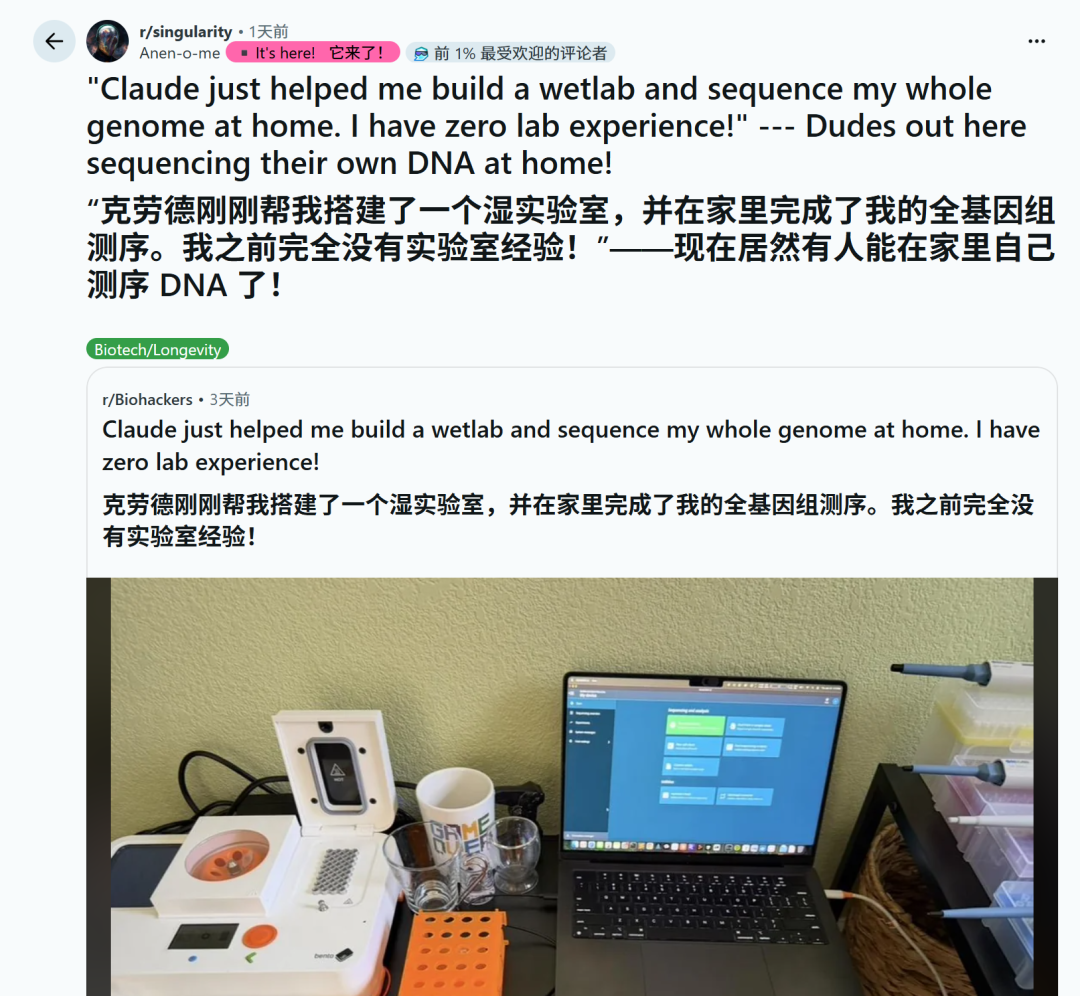
Laboratory on table, including heating module, vortexer, mini centrifuge, pipette, MinION
Now, this post has hit the internet.
People who have done experiments in the laboratory for ten years say with emotion: You are so professional.
People who do genetic research say that this genome sequencing project fits their research scope.
A patient suffering from tumors said that this experiment is of great significance.

It can be said that this is a DIY practice that can shock the entire biosphere: from then on, biological research broke the monopoly of institutions and completely entered the era of individuals!
He hopes to regain the decision-making power of life from the pain
Seth Howes is a former exolabs engineer and holds an MD from Oxford and a PhD in machine learning from Imperial College.
Seth's research did not originate from academic interest, but from a heavy family genetic pressure.
His family has a long history of high-risk autoimmune diseases.
While he was conducting experiments, his sister, who was less than 40 years old, suffered severe liver damage due to illness. She had to wait two years before obtaining a liver source for surgery.

“I have no illusions that we can cure our family’s ills, but I do want to understand why our bodies continue to eat themselves back, generation after generation.”
It was this curiosity about the underlying code of life, coupled with the pursuit of data privacy, that prompted him to decide to build a "digital apothecary" workbench in his living room.
After all, in his opinion, "your most private data should never leave your house."
Core black technology: exponential blessing of AI
In the past, whole-genome sequencing was the preserve of large laboratories and cost hundreds of thousands of dollars.
The key to Seth's success this time lies in the blessing of three elements.
Hardware: A "gene reader" in your pocket
First, in terms of hardware, he used the Oxford Nanopore MinION sequencer.
This thing is only the size of a USB flash drive, and it can read DNA sequences when plugged into a computer.
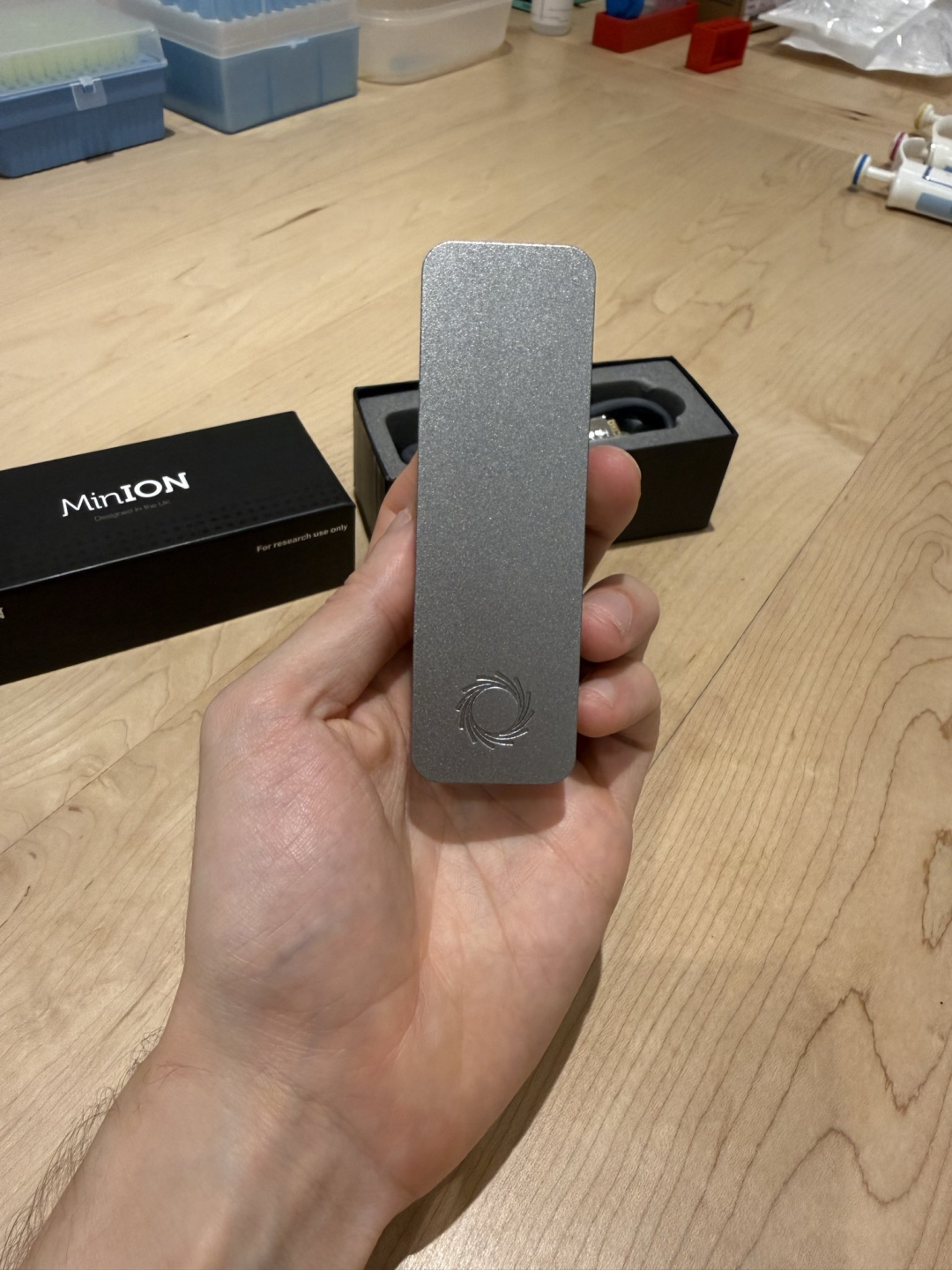

A few years ago, the same thing would have required a room-filling Illumina sequencer, a dedicated team, and a six-figure budget.
Now, this device, which is only the size of a power bank, is filled with about 2,000 nanopores (only 1 nanometer in diameter). When DNA fragments pass through these holes, the microcurrent changes caused will be recorded and converted into genetic codes.
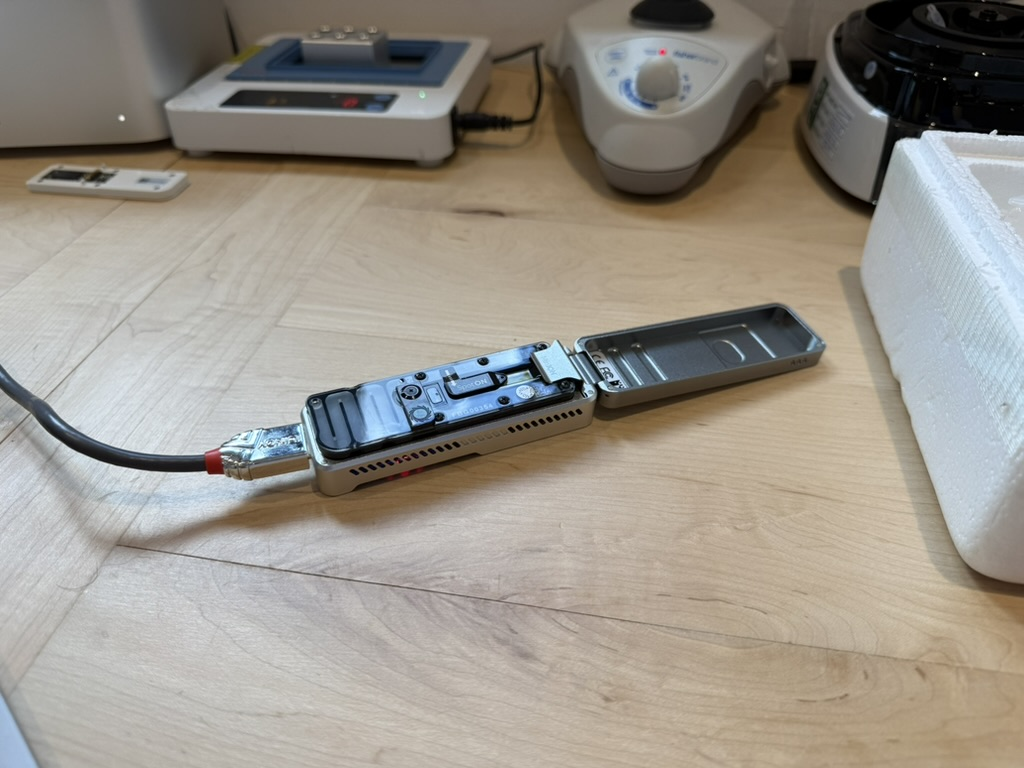

It reduces the cost of sequencing from hundreds of thousands of dollars to the $1,000 level, and may even drop to $100 in the future.
It can be said that MinION has transformed "reading DNA" from a capital-intensive activity into a tool-based capability.
Just like 3D printers move "manufacturing" from the factory to the desktop, MinION moves "sequencing" from the laboratory to the living room.
AI model: from "reading code" to "understanding functions"
But in the experiment, it is not enough to read the four bases A, T, C, and G.
Three billion base pairs of raw data are in front of you, and if you don't know what they mean, it's just a bunch of letters.
This is where AI comes into play. In the experiments, two key models were used.
The first one is Evo2.
This is a basic genome model developed by the Arc Institute, with a parameter scale of 7 billion and trained on the genome data of more than 120,000 species around the world. What it can do is: given a DNA sequence, predict the biological function of this sequence.
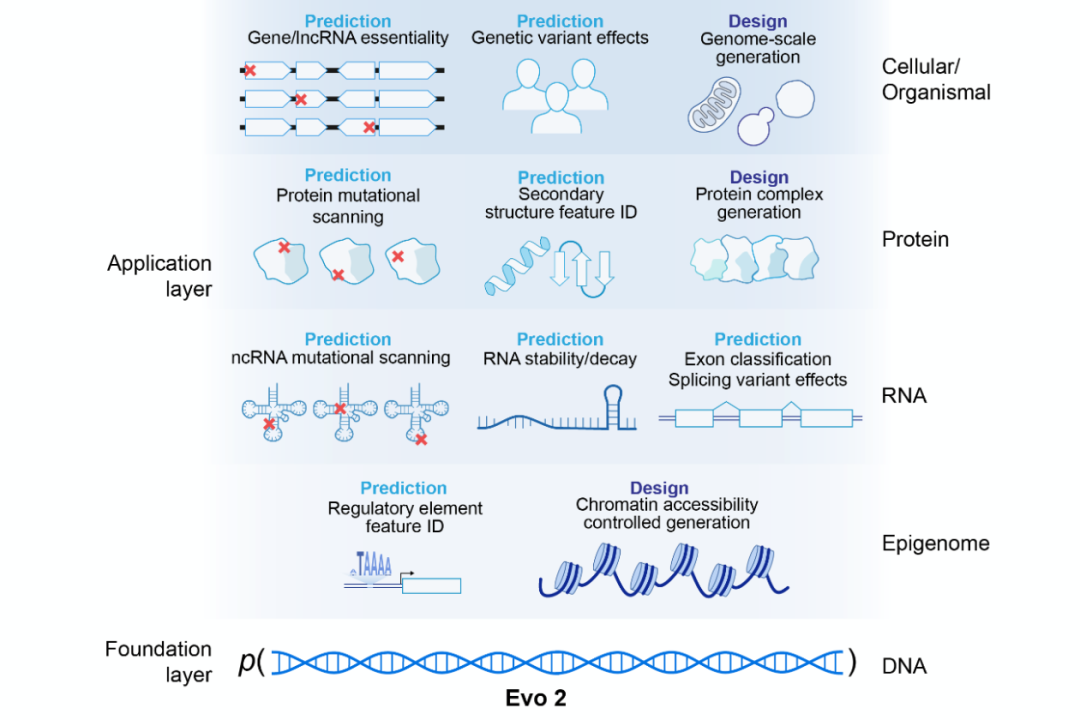
You must have realized it - this is a biological version of GPT!
GPT understands human language, and Evo2 understands the language of life. The difference is, the book Evo2 reads is 3 billion letters long.
The second one is AlphaGenome.
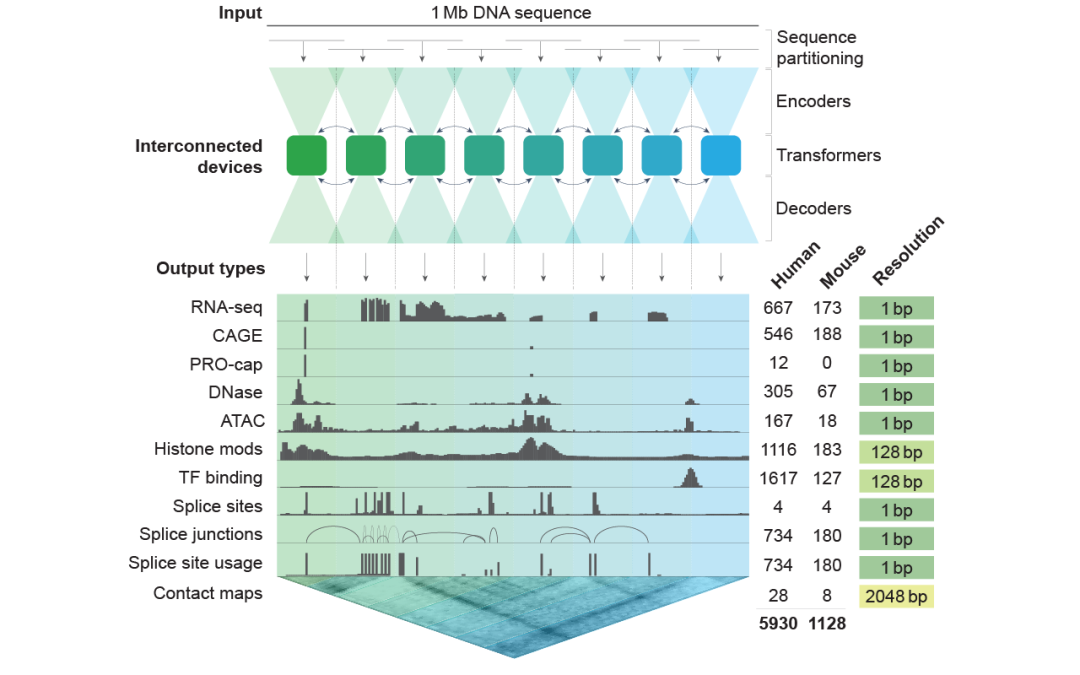
Produced by Google DeepMind, which specializes in genome function prediction. It not only tells you "what protein is encoded by this DNA", but also predicts "what impact a mutation at this site will have on gene expression and chromatin structure."
From "reading DNA" to "understanding DNA function", this leap used to take an entire molecular biology laboratory several months to verify. Now, a model can run for just a few hours and the results will be available.
Simplified interface: large models become "laboratory assistants"
Speaking of which, the most surprising detail in the whole thing was not the sequencer or the genome model, but Claude.
Seth used Claude to generate the BED file during the operation.
BED files are a standard data format for genomics, recording the coordinate information of specific regions on the genome. In the past, writing this kind of file required professionals who knew bioinformatics to write it manually or use specialized command line tools.
The current approach is to use natural language to tell Claude "generate a BED file for me to cover these gene regions", and it will be generated.
Biological operations,have been taken over by language interfaces.
The era of personal laboratories is here
Now, the democratization of hardware (MinION), the exponential improvement of AI understanding (Evo2 + AlphaGenome), and the smoothing of the operating threshold by language interfaces (Claude) directly constitute a complete paradigm shift chain.
Each link alone is not news. But when the three rings close at the same time, something goes wrong!
A curious young man solved a mystery in his living room that had eluded clinicians for more than ten years.
The impact of this incident is not that Seth’s personal story is legendary, but that it reveals an accelerating curve.
The downward trajectory of sequencing costs is even more severe than Moore’s Law.
In 2003, the cost of completing a human genome sequence was $2.7 billion. In 2007 it dropped to 10 million. It dropped to $1,000 in 2014. In 2024, some platforms will already be able to achieve prices below US$200.
The next goal is $100.

The slope of this curve means that in the not-too-distant future, sequencing your own genome may cost less than a full physical exam.
Many people’s first reaction is: Isn’t this just a hobby project for geeks? Who among ordinary people would test their genome at home?
But think back, in 2010, how many people thought that “ordinary people would use 3D printers at home”?
Every time the tool chain is closed, the window period from geek toys to mass applications is shortened. The window period may be shorter this time because the driving force behind it is rigid demand.
Living room experiment record: How to "read" your own DNA at home?
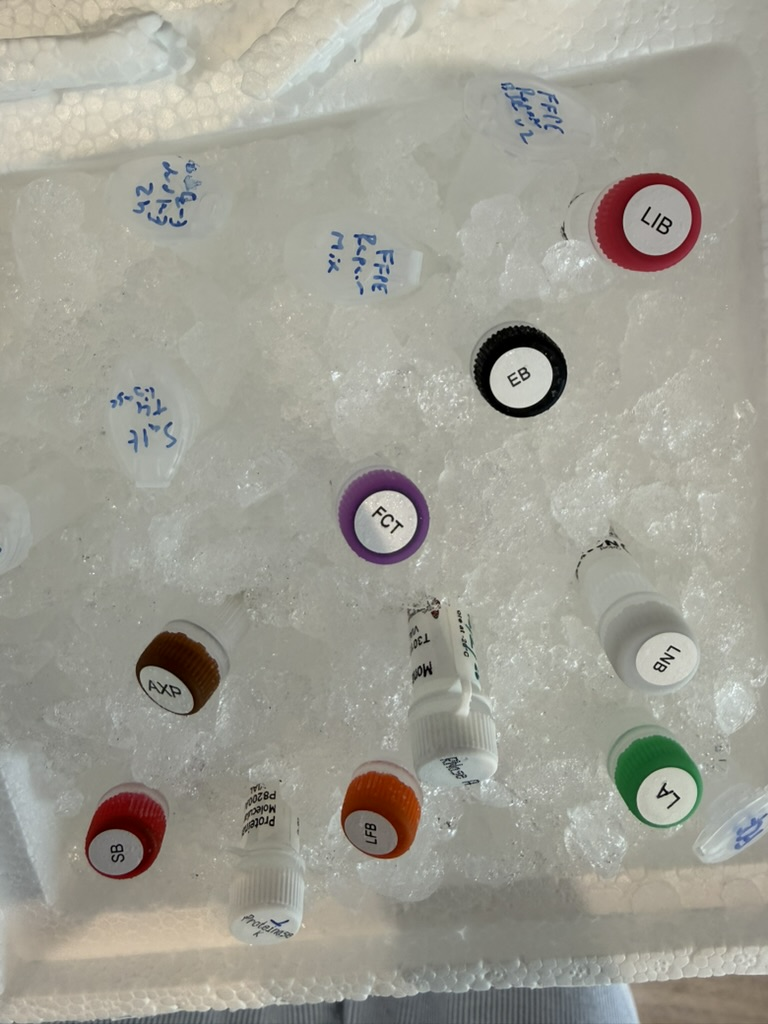
The complete operating procedures published by Seth show that in addition to precision pipettes, many of his laboratory equipment are second-hand from eBay or AliExpress.
In the sample extraction stage, he did not take complicated blood draws, but rubbed the inside of his cheek with a sterile oral swab. He obtained about 5-7 micrograms of DNA with only 1 swab, far exceeding the 1 microgram required for the experiment.
Adaptive sampling is MinION’s trump card.
Seth uses software control to let the sequencer perform a "preliminary screening" when reading the first 500 bases of each fragment: if it is an immune-related gene he is concerned about, continue reading; if not, reverse the voltage to "spit out" the DNA strand and replace it with the next one.
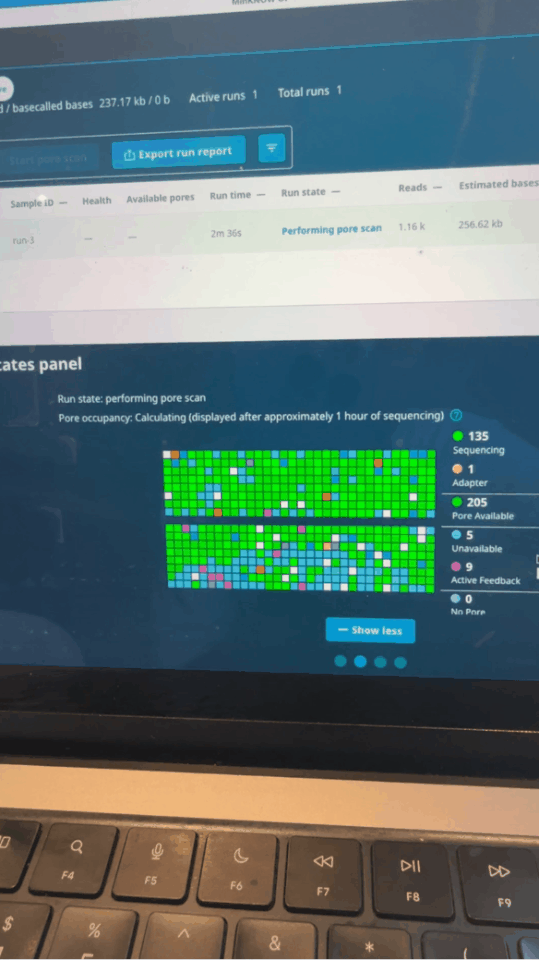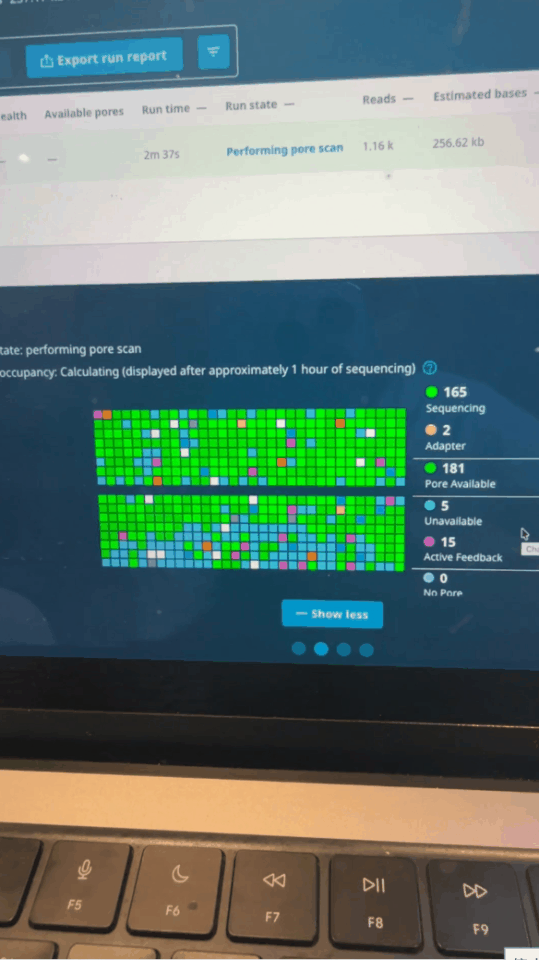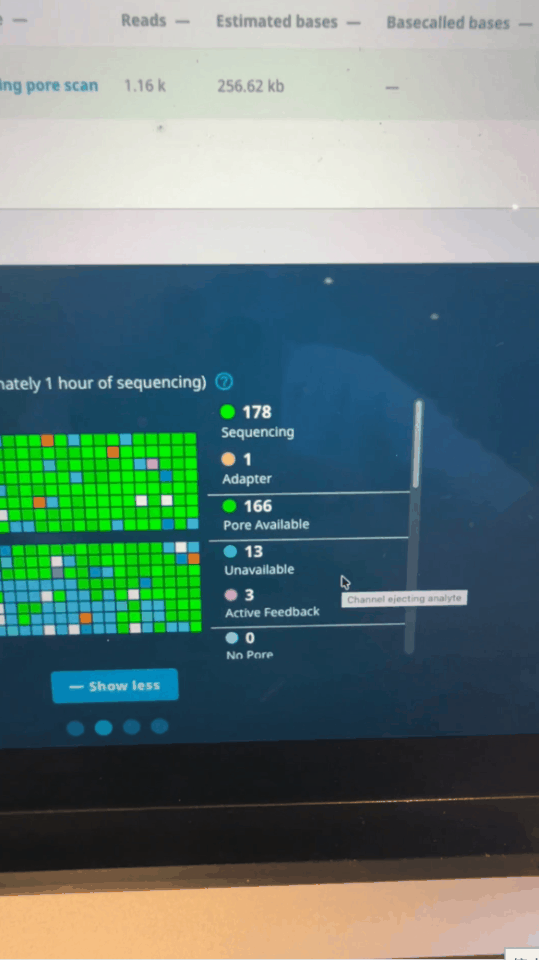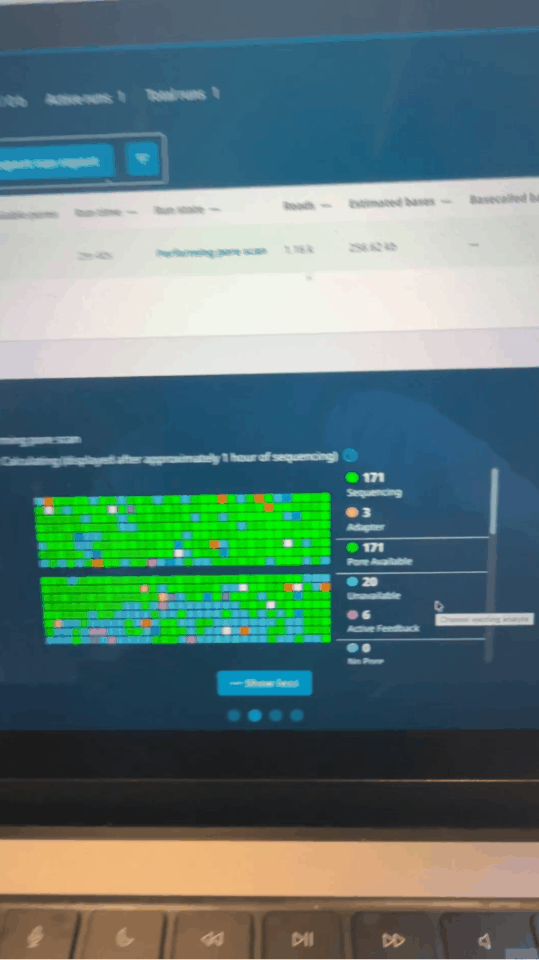
Nanopore activity status on MinKNOW control panel minutes after run starts
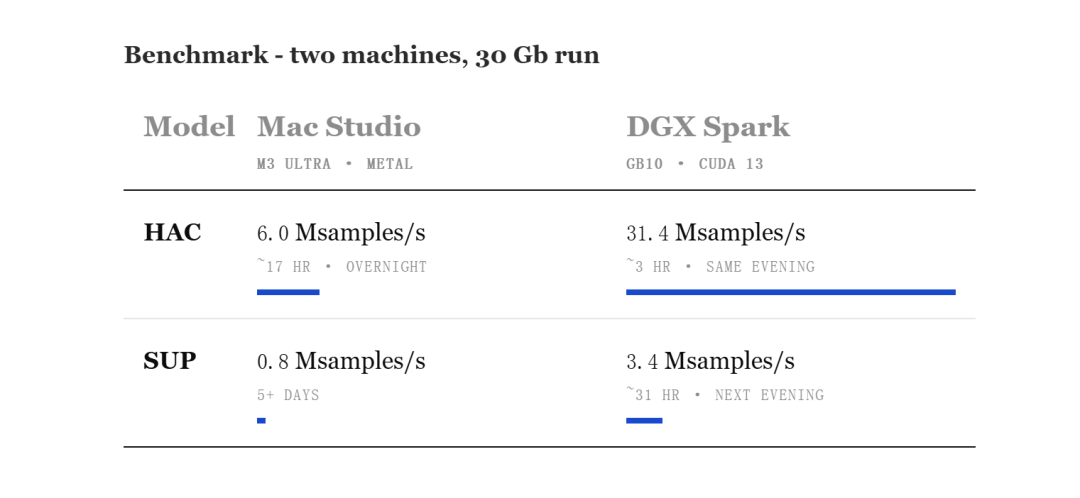
In this process, he used the M3 Ultra chip's Mac Studio for real-time base identification, and used NVIDIA's DGX Spark to increase the analysis speed and ensure computing power.
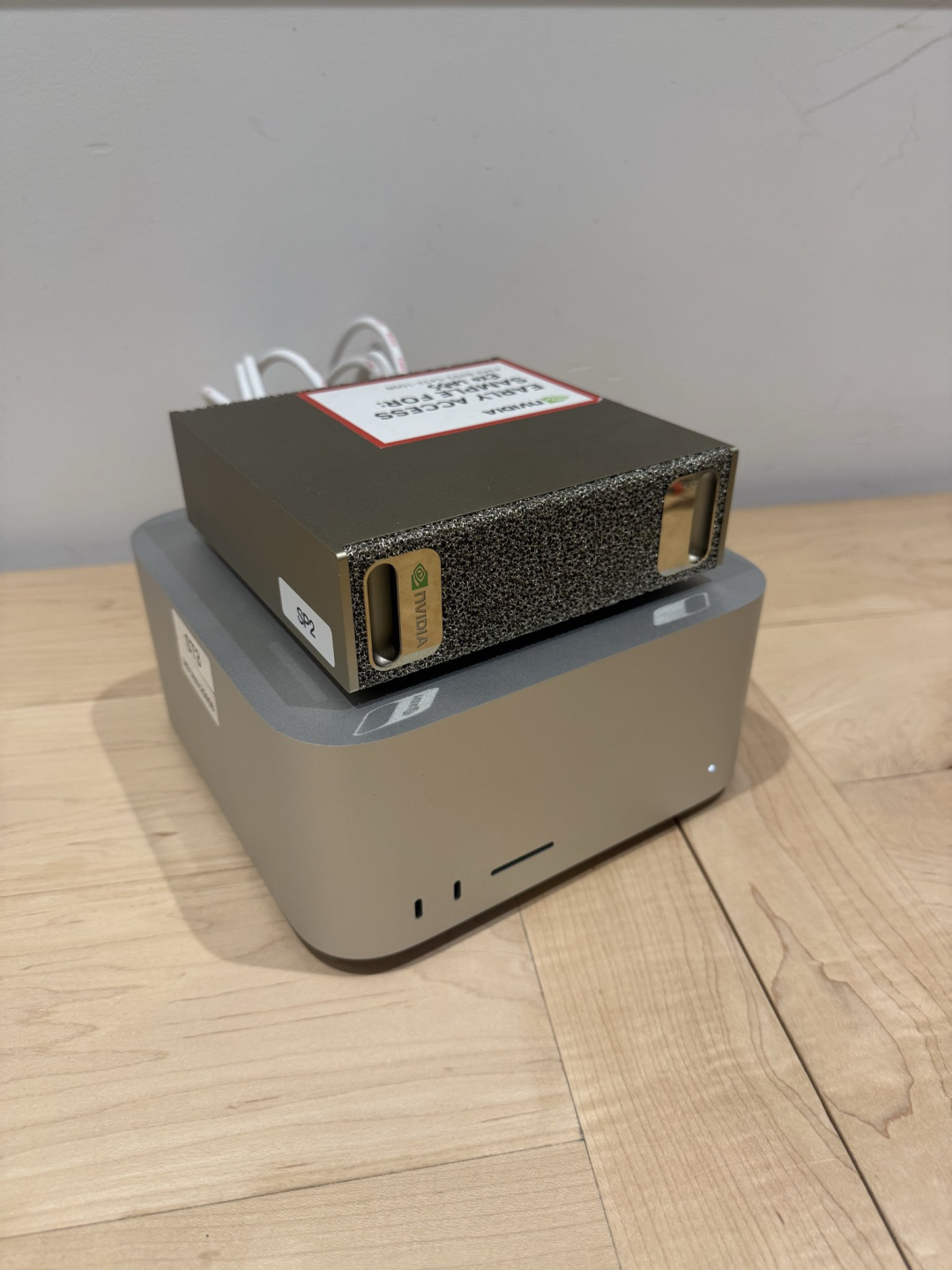

Biology is no longer an individual monopoly
In the 1970s, computers were the preserve of large organizations, then personal computers came along, and then the Internet came along.
Biology is going the same way.
The sequencer is the personal computer, the genome base model is the operating system, and Claude is the user interface. When all three layers are in place simultaneously, biology's "PC moment" will arrive.
The afternoon Seth finished sequencing his genome in his living room may have been the first frame of this moment.
Looking back ten years later, 2026 will probably be a watershed year.
The significance of this experiment goes far beyond the victory of a tech geek. It marks that biological research is undergoing a paradigm shift from institutional monopoly to individual DIY mode.
When complex biological operations are simplified into language conversations by AI, and when expensive instruments become affordable for everyone, ordinary people will be able to actively analyze the underlying logic of life.
As Seth said, he found the pleasure of playing with computers when he was a child when playing with genetic circuits. This "geek spirit" is opening a new era.
How many more miracles will AI create next?